Comprehensive molecular analysis of several prognostic signatures using molecular indices... to hallmarks of breast cancer: proliferation index appears to be...

Survival Analysis DMFS (original data sets)
Conclusions
Proliferation seems to be the
common denominator of many
existing prognostic gene
signatures, recapitulating their
prognostic power.
Printed by
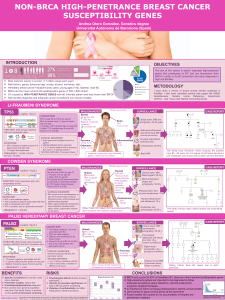
Comprehensive molecular analysis of several prognostic signatures using molecular indices related
to hallmarks of breast cancer: proliferation index appears to be the most significant component of all signatures
Christos Sotiriou1,2, Benjamin Haibe-Kains1,2,3, Christine Desmedt1,2, Pratyaksha Wirapati4, Virginie Durbecq1, Adrian Harris5, Denis Larsimont1,
Gianluca Bontempi3, Marc Buyse6, Mauro Delorenzi4, and Martine Piccart1,3.
1Jules Bordet Institute, 2National Foundation of Cancer Research,3Free University of Brussels, 4Swiss Institute of Bioinformatics, Lausanne, 5John Radcliff Hospital, Oxford, UK, 6IDDI,Brussels
Defining Molecular Indices
(Sotiriou et al. ASCO 2006)
Representative genes
Van de Vijver
et al.
NEJM, 2002
2 Published datasets
Agilent
N=295
Wang
et al.
Lancet, 2005
Affymetrix
N=251
ESR1 = Basal/luminal
ERBB2 = Her2-neu
STK6 = proliferation/GGI
PLAU = Stroma/invasion
STAT1 = immune response
gene X1
gene X40000
.
.
.
.
+
+
+
+
A model selection procedure is fitted to
estimate the contribution of each
prototype for the prediction of the
expression of each gene on the arrays CASP3 = apoptosis
+
VEGF = angiogenesis
+
70 genes
Van’t veer et
al. Nature
2002
76 genes
Wang et al.
Lancet 2005
P53
(32 genes)
Miller at al.
PNAS 2005
Wound
(573 genes)
Chang et al.
Plos Biology
2004
Genomic grade
(97 genes)
Sotiriou et al.
JNCI 2006
Recurence
Score
(21 genes)
Pail et al.
NEJM 2004
ESR1
(lumina/basal)
73%
(10%)
38%
(3%)
88%
(34%)
42%
(4%)
73%
(1%)
37%
(2%)
99%
(54%)
64%
(0%)
43%
(0%)
STAT1
(immune response)
29%
(1%)
30%
(0%)
19%
(3%)
30%
(0%)
43%
(0%)
25%
(0%)
30%
(0%)
69%
(19%)
ERBB2 60%
(0%)
35%
(0%)
53%
(0%)
30%
(0%)
44%
(6%)
STK6
(proliferation)
63%
(14%)
55%
(16%)
53%
(16%)
52%
(13%)
69%
(13%)
PLAU
(stroma/invasion)
47%
(3%)
42%
(5%)
47%
(0%)
39%
(3%)
38%
(6%)
VEGF
(angiogenesis)
43%
(0%)
26%
(1%)
28%
(0%)
35%
(1%)
25%
(0%)
CASP3
(apoptosis)
60%
(0%)
16%
(1%)
38%
(0%)
40%
(3%)
38%
(0%)
Dissecting Gene Expression
Prognostic Signatures
% of genes related to each prototype; % of genes specific for each prototype
Introduction
•Selection of prototype genes related to several biological processes in
breast cancer (hallmarks of cancer) such as basal/luminal phenotype,
ERBB2, proliferation, fully captured by the gene expression grade index,
stroma/invasion, angiogenesis, apoptosis and immune response (Sotiriou et
al ASCO 2006)
Although the development of high-throughput gene expression
technologies has led to the identification of several “molecular
signatures” predicting clinical outcome, no attempt has yet been made
to perform a comprehensive analysis integrating well characterized
biological processes and gene expression data. Here we aim to
elucidate the relationship of gene expression patterns defined by
several biologically relevant indices with previously reported prognostic
signatures and their interaction with prognosis
Materials & Methods
•Use of a model selection procedure based on cross-validation error
estimation in order to select the genes that are able to predict
significantly et specifically one of the prototypes (molecular indices)
•Application to several previously reported prognostic signatures (70-
gene, 76-gene, wound healing, p53, genomic grade and recurrence score)
•Characterization of dependency patterns between these indices for
each prognostic signature and their impact on survival using several
microarray datasets
Hallmarks of breast cancer
Hanahan D, Weinberg R, Cell 2000
HER-2
ER/luminal/bas
al
uPA/PAI
Immune
Response
Proliferation
genes (GGI)
VEGF
Survival Analysis DMFS, TRANSBIG VALIDATION N=198
Van’t veer et al. Wang et al. Sotiriou et al.
Acknowledgments
FNRS, Medic Foundation,
Breast Cancer Research Foundation
(Evelyne Lauder)
Van’t veer et al. Nature 2002, 70 genes
Wang et al. Lancet 2005, 76 genes
Miller at al. PNAS 2005, p53, 32 genes
Chang et al. Plos Biology 2004, wound, 573 genes
Sotiriou et al. JNCI 2006, genomic grade, 97 genes
Paik et al. NEJM 2004, recurence score, 21 genes
1
/
1
100%
![[PDF]](http://s1.studylibfr.com/store/data/008642620_1-fb1e001169026d88c242b9b72a76c393-300x300.png)